Introduction: The Next Frontier After Protein Folding
The 2024 Nobel Prize awarded to AlphaFold2 underscored the transformative role of artificial intelligence in biology. Predicting protein structures from sequences was a monumental achievement, but the field now asks: what comes next? The answer lies in generation—not just predicting existing proteins, but designing new ones with desired functions. PLAID (Protein Latent diffusion for AI-based Design) is a groundbreaking model that repurposes the latent space of protein folding models to generate novel proteins. Unlike prior work, PLAID tackles the complex task of multimodal co-generation: simultaneously producing both the discrete amino acid sequence and the continuous all-atom three-dimensional structure.
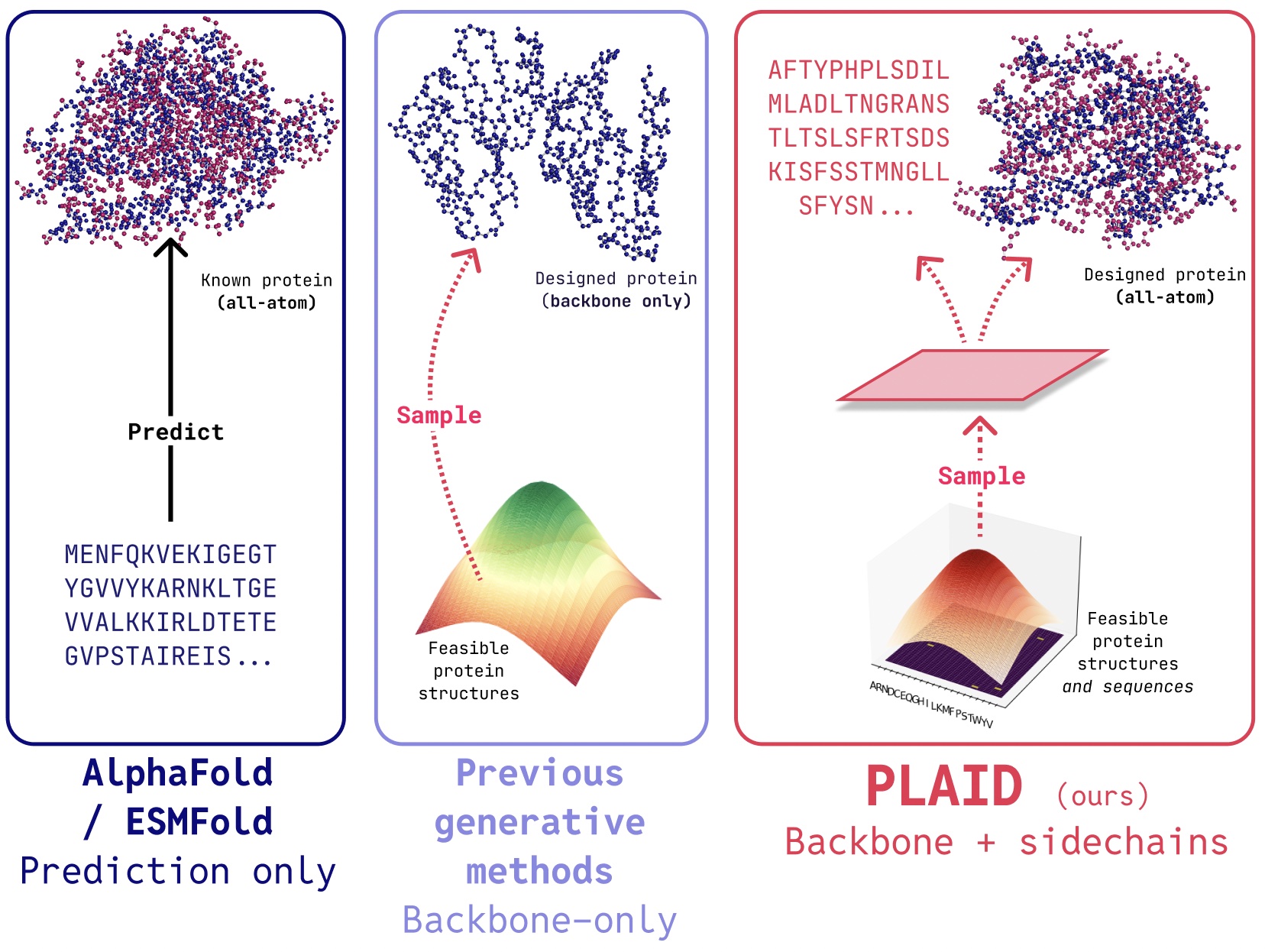
From Structure Prediction to Protein Design
While diffusion models have shown promise for protein generation, earlier approaches faced practical limitations that prevented real-world deployment:
- All-atom generation: Many existing models generate only backbone atoms (Cα, C, N). To place side-chain atoms, the sequence must be known, creating a chicken-and-egg problem that demands simultaneous generation of discrete (sequence) and continuous (structure) modalities.
- Organism specificity: Protein biologics intended for human use must be “humanized” to evade the immune system. A generative model must respect the target organism’s context.
- Control specification: Drug discovery requires navigating complex constraints—e.g., solubility for tablet formulation versus vial transport, or binding affinity to a specific receptor. How do we encode such multifaceted requirements into a generative process?
PLAID directly addresses these challenges by learning a latent space that captures the relationship between sequence, structure, and function, enabling controlled generation through compositional prompts.
PLAID's Multimodal Generation Framework
At its core, PLAID learns to sample from the latent space of protein folding models (such as those used in AlphaFold) to generate new protein sequences and structures simultaneously. This is achieved via a latent diffusion process that operates on a joint representation of sequence and structure. Key innovations include:
- Compositional prompts: Users can specify functional annotations (e.g., “metal-binding” or “kinase”) and target organism (e.g., “human” or “E. coli”) as textual constraints. The model then generates proteins that satisfy these conditions.
- Sequence-only training: While structure databases are limited (∼200,000 entries), sequence databases contain millions to billions of sequences. PLAID can be trained exclusively on sequences, leveraging vastly larger datasets to learn the sequence–structure–function mapping.
- All-atom output: The model produces complete all-atom coordinates including side chains, eliminating the need for post-processing.
Generating Useful Proteins with Controllable Interfaces
Simply generating random proteins is not useful; control is essential. PLAID’s interface draws inspiration from image generation, where compositional textual prompts (e.g., “a red car on a snowy road”) allow fine-grained control (Liu et al., 2022). In PLAID, the analogous prompts use two axes: function and organism. For example, prompting for a “humanized metalloprotein with zinc-binding activity” would constrain generation to proteins that are structurally compatible with human biology and carry a specific catalytic function. This paves the way for designing therapeutics, enzymes, and biosensors with predefined properties.
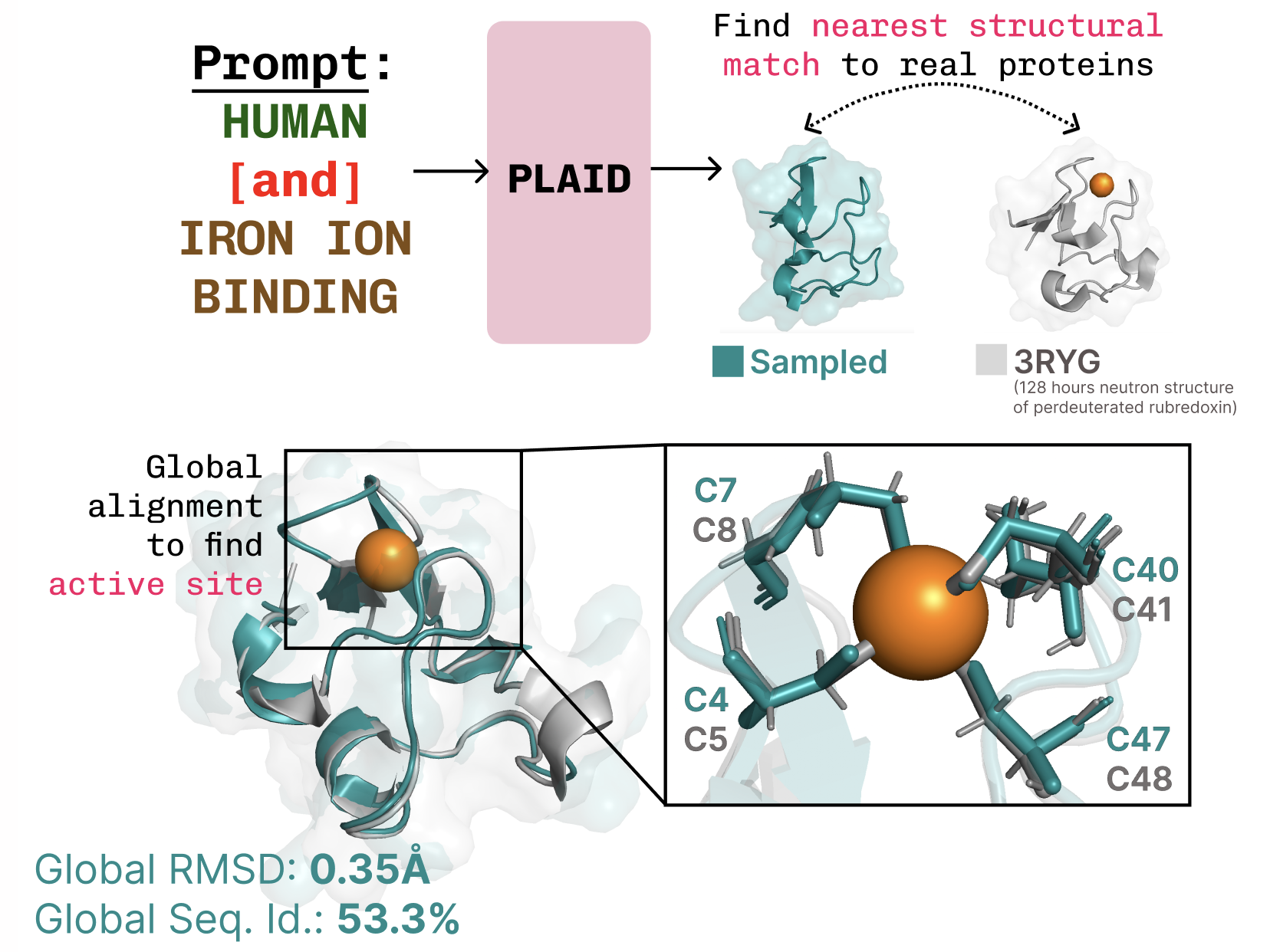
As an earlier noted, organism specificity is critical for biologics. PLAID’s organism prompt can be trained on sequence data from any species, enabling humanization or even cross-species design.
Learning the Function-Structure-Sequence Connection
A striking example of PLAID’s capability is its learning of the tetrahedral cysteine-Fe2+/Fe3+ coordination pattern—a motif common in metalloproteins like cytochromes and iron-sulfur clusters. Despite being trained only on sequences, the latent diffusion model reproduces this geometric arrangement with high fidelity, while maintaining high sequence-level diversity. This shows that PLAID internalizes the structural constraints encoded in sequence space, without ever seeing explicit 3D coordinates during training.
Training with Sequence-Only Data: A Game Changer
One of PLAID’s most practical advantages is its ability to be trained using only sequence databases, which are 2–4 orders of magnitude larger than structure databases. Sequences are inexpensive to obtain (via gene sequencing), while experimental structures require costly X-ray crystallography, NMR, or cryo-EM. By learning the generative distribution from sequences alone, PLAID can scale to millions of protein families, capturing evolutionary diversity and functional patterns that structure-based models might miss. This approach mirrors the success of large language models in NLP, where massive text corpora enable rich representations.
Conclusion: Toward Real-World Protein Design
PLAID represents a significant stride beyond structure prediction into active protein design. Its latent diffusion framework, compositional prompts, and sequence-only training make it a versatile tool for generating functional proteins with controlled properties. As the field moves toward practical applications—from enzyme engineering to therapeutic antibody design—models like PLAID will be essential for navigating the vast sequence–structure–function landscape. The future of protein design is here, and it is generative.